FIELD RESEARCH
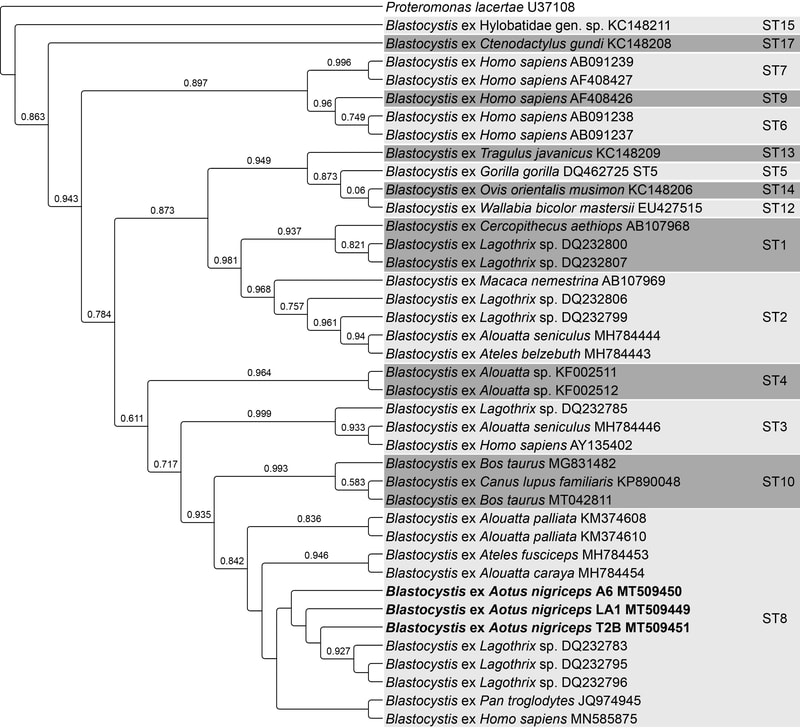
Phylogenomics, evolutionary history, and taxonomy
The classification and evolutionary history of night monkeys has continued to be repeatedly challenged using cytogenetic, morphological, and molecular data, resulting in the ongoing revision of taxonomic and evolutionary history. This research applies expanded phylogenomics to assess diversity of the genus. The implications of revised taxonomy are wide-ranging. Several Aotus taxa are found in areas with the highest deforestation rates in the world driven by growing impacts from fire related to agricultural expansion into primary tropical rainforest in the Amazon.
Night Monkey Behavior and Ecology
One of our main projects is working with the black-headed night monkey, Aotus nigriceps. Night monkeys - also known as owl monkeys - are the only group of true nocturnal monkey species. Since they are nocturnal, researchers traditionally have avoided studying them due to the logistical difficulty of working at night. Thus, information on Aotus behavior, ranging patterns, habitat, ecology, and conservation status is generally limited compared to many other primates. We're here to change that!
Biogeography and Conservation
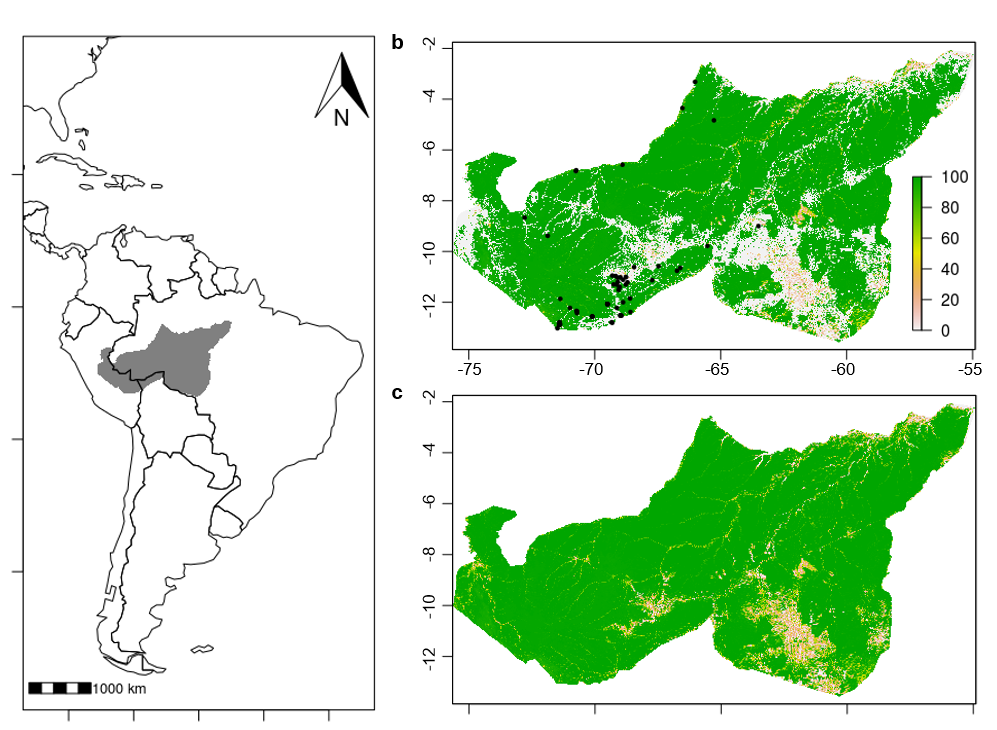
Primate populations are declining worldwide as their habitat is degraded by expanding anthropogenic disturbances. It's still unclear how Aotus reacts to habitat disturbance. We are analyzing habitat preferences including the use of bamboo species, establishing population level density estimates, and assessing diet composition across seasons and habitat types. We are also investigating population structure of A. nigriceps throughout their range.
Parasitology
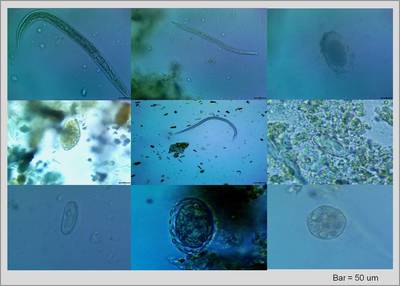
Gastrointestinal parasites are a main driver of mortality, morbidity, and mating/group behavior in wild non-human primate groups. Several lines of thought have suggested that gastrointestinal parasite species richness may be increasing with anthropogenic disturbance. We have recovered a numerous intestinal parasites including Strongyloides spp. and hookworm spp., along with the less common Trypanoxyuris sp., Physaloptera sp., Ascarid sp., Entamoeba sp., Enterobius sp. and Trichostrongylus sp. We continue to study the impact of forest type and disturbance on intestinal parasitism.
Acoustics
Our objective is to gain a better understanding of Aotus nigriceps vocalization diversity, context, group and population differences, and observe the behavior and auditory responses to predators and competitors. We have described new call characteristics, and several differences in call frequencies have been observed between groups. We continue to study differences and similarities between groups.
Phylogenomics, evolutionary history, and taxonomy
The classification and evolutionary history of night monkeys has continued to be repeatedly challenged using cytogenetic, morphological, and molecular data, resulting in the ongoing revision of taxonomic and evolutionary history. This research applies expanded phylogenomics to assess diversity of the genus. The implications of revised taxonomy are wide-ranging. Several Aotus taxa are found in areas with the highest deforestation rates in the world driven by growing impacts from fire related to agricultural expansion into primary tropical rainforest in the Amazon.
Night Monkey Behavior and Ecology
One of our main projects is working with the black-headed night monkey, Aotus nigriceps. Night monkeys - also known as owl monkeys - are the only group of true nocturnal monkey species. Since they are nocturnal, researchers traditionally have avoided studying them due to the logistical difficulty of working at night. Thus, information on Aotus behavior, ranging patterns, habitat, ecology, and conservation status is generally limited compared to many other primates. We're here to change that!
Biogeography and Conservation
Primate populations are declining worldwide as their habitat is degraded by expanding anthropogenic disturbances. It's still unclear how Aotus reacts to habitat disturbance. We are analyzing habitat preferences including the use of bamboo species, establishing population level density estimates, and assessing diet composition across seasons and habitat types. We are also investigating population structure of A. nigriceps throughout their range.
Parasitology
Gastrointestinal parasites are a main driver of mortality, morbidity, and mating/group behavior in wild non-human primate groups. Several lines of thought have suggested that gastrointestinal parasite species richness may be increasing with anthropogenic disturbance. We have recovered a numerous intestinal parasites including Strongyloides spp. and hookworm spp., along with the less common Trypanoxyuris sp., Physaloptera sp., Ascarid sp., Entamoeba sp., Enterobius sp. and Trichostrongylus sp. We continue to study the impact of forest type and disturbance on intestinal parasitism.
Acoustics
Our objective is to gain a better understanding of Aotus nigriceps vocalization diversity, context, group and population differences, and observe the behavior and auditory responses to predators and competitors. We have described new call characteristics, and several differences in call frequencies have been observed between groups. We continue to study differences and similarities between groups.
Publications (ORCID ID)
Helenbrook, W. D., & Valdez, J. (Preprint: 2023). The role of rivers as geographical barriers in shaping genetic differentiation and diversity of Neotropical primates. bioRxiv, 2023-07.
Helenbrook, W.D. and Valdez, J. (2021). Species distribution and conservation assessment of the black-headed night monkey (Aotus nigriceps): a species of Least Concern that faces widespread anthropogenic threats. Primates
Helenbrook, W.D. and Whipps, C. (2021). Molecular characterization of Blastocystis in captive and free-ranging New World Primates. Acta Parasitologica.
Helenbrook, W.D. and Valdez, J. (2020). Species distribution modeling and conservation assessment of the black-headed night monkey (Aotus nigriceps) – A species of Least Concern that faces widespread anthropogenic threats. PrePrint bioRxiv
Helenbrook, W.D., Nelson, A., and Solorzano, B. (2020). Intestinal parasitism in the black-headed night monkey, Aotus nigriceps. International Journal of Primatology.
Helenbrook, W.D., Wilkinson, M., and Suarez, J. (2020). Habitat use, fruit consumption, and population density of the black-headed night monkey, Aotus nigriceps, in Southeastern Peru. Acta Amazonica.
Helenbrook, W.D., Linck, N., Pardo, M., and Suarez, J. (2019). Spatial variation in black headed night monkey (Aotus nigriceps) vocalizations. PrePrint bioRxiv
Helenbrook, W.D., Preston, L., Linck, N., Quirk, M., and Suarez, J. (2018). Bioacoustics of the black-headed night monkey, Aotus nigriceps. Neotropical Primates 24: 29-33
Helenbrook, W.D., Stehman, S.V., Shields, W.M., & Whipps, C.M. (2017). Association of anthropogenic disturbances and intestinal parasitism in Ecuadorian mantled howler monkeys, Alouatta palliata aequatorialis. Folia Primatologica 88: 307-322.
Helenbrook, W. D., Shields, W. M., & Whipps, C. M. (2015). Characterization of Blastocystis species infection in humans and mantled howler monkeys, Alouatta palliata aequatorialis, living in close proximity to one another. Parasitology Research 114: 2517-2525.
Helenbrook, W.D., Wade, S.E., Shields, W.M., Stehman, S.V., & Whipps, C.M. (2015). Gastrointestinal parasites of Ecuadorian mantled howler monkeys (Alouatta palliata aequatorialis) based on fecal analysis. The Journal of parasitology 101: 341-350.
Helenbrook, W. (2015). Evaluation of mandrill monkey fecal preservation and extraction methodologies reveal ideal combinations for maximum DNA yields. Report. Tropical Conservation Fund.
Helenbrook, W.D. (2014). Effects of ecological disturbance on parasite communities in both people and mantled howler monkeys (Alouatta palliata aequatorialis) living in Ecuador. State University of New York College of Environmental Science and Forestry.
Helenbrook, W.D. (2014). Risk factors associated with endoparasitism in two rural Ecuadorian communities. State University of New York College of Environmental Science and Forestry.
Searing, K.B. , Shoemaker, K.T., Helenbrook, W.D., Schlaepfer, M.A. (2010). Move It? An assessment tool and ranking system for assisted colonization. PS 56-96: Move It? An assessment tool and ranking system for assisted colonization. In The 95th ESA Annual Meeting.
Schlaepfer, M.A., Helenbrook, W. D., Searing, K. B., & Shoemaker, K. T. (2009). Assisted colonization: evaluating contrasting management actions (and values) in the face of uncertainty. Trends in Ecology & Evolution, 24(9), 471-472.
Helenbrook, W.D. (2006). Non-invasive sampling of mandrill and drill monkeys for use in genetic analyses. In American Journal of Primatology 68, pp. 88-88).
Helenbrook, W. D., & Valdez, J. (Preprint: 2023). The role of rivers as geographical barriers in shaping genetic differentiation and diversity of Neotropical primates. bioRxiv, 2023-07.
Helenbrook, W.D. and Valdez, J. (2021). Species distribution and conservation assessment of the black-headed night monkey (Aotus nigriceps): a species of Least Concern that faces widespread anthropogenic threats. Primates
Helenbrook, W.D. and Whipps, C. (2021). Molecular characterization of Blastocystis in captive and free-ranging New World Primates. Acta Parasitologica.
Helenbrook, W.D. and Valdez, J. (2020). Species distribution modeling and conservation assessment of the black-headed night monkey (Aotus nigriceps) – A species of Least Concern that faces widespread anthropogenic threats. PrePrint bioRxiv
Helenbrook, W.D., Nelson, A., and Solorzano, B. (2020). Intestinal parasitism in the black-headed night monkey, Aotus nigriceps. International Journal of Primatology.
Helenbrook, W.D., Wilkinson, M., and Suarez, J. (2020). Habitat use, fruit consumption, and population density of the black-headed night monkey, Aotus nigriceps, in Southeastern Peru. Acta Amazonica.
Helenbrook, W.D., Linck, N., Pardo, M., and Suarez, J. (2019). Spatial variation in black headed night monkey (Aotus nigriceps) vocalizations. PrePrint bioRxiv
Helenbrook, W.D., Preston, L., Linck, N., Quirk, M., and Suarez, J. (2018). Bioacoustics of the black-headed night monkey, Aotus nigriceps. Neotropical Primates 24: 29-33
Helenbrook, W.D., Stehman, S.V., Shields, W.M., & Whipps, C.M. (2017). Association of anthropogenic disturbances and intestinal parasitism in Ecuadorian mantled howler monkeys, Alouatta palliata aequatorialis. Folia Primatologica 88: 307-322.
Helenbrook, W. D., Shields, W. M., & Whipps, C. M. (2015). Characterization of Blastocystis species infection in humans and mantled howler monkeys, Alouatta palliata aequatorialis, living in close proximity to one another. Parasitology Research 114: 2517-2525.
Helenbrook, W.D., Wade, S.E., Shields, W.M., Stehman, S.V., & Whipps, C.M. (2015). Gastrointestinal parasites of Ecuadorian mantled howler monkeys (Alouatta palliata aequatorialis) based on fecal analysis. The Journal of parasitology 101: 341-350.
Helenbrook, W. (2015). Evaluation of mandrill monkey fecal preservation and extraction methodologies reveal ideal combinations for maximum DNA yields. Report. Tropical Conservation Fund.
Helenbrook, W.D. (2014). Effects of ecological disturbance on parasite communities in both people and mantled howler monkeys (Alouatta palliata aequatorialis) living in Ecuador. State University of New York College of Environmental Science and Forestry.
Helenbrook, W.D. (2014). Risk factors associated with endoparasitism in two rural Ecuadorian communities. State University of New York College of Environmental Science and Forestry.
Searing, K.B. , Shoemaker, K.T., Helenbrook, W.D., Schlaepfer, M.A. (2010). Move It? An assessment tool and ranking system for assisted colonization. PS 56-96: Move It? An assessment tool and ranking system for assisted colonization. In The 95th ESA Annual Meeting.
Schlaepfer, M.A., Helenbrook, W. D., Searing, K. B., & Shoemaker, K. T. (2009). Assisted colonization: evaluating contrasting management actions (and values) in the face of uncertainty. Trends in Ecology & Evolution, 24(9), 471-472.
Helenbrook, W.D. (2006). Non-invasive sampling of mandrill and drill monkeys for use in genetic analyses. In American Journal of Primatology 68, pp. 88-88).